- Applications
-
Products
-
Liquid Handling
- firefly Accelerate genomic research with innovative all-in-one, compact liquid handling
- mosquito Nanolitre liquid handling technology performs ‘traditional’ tasks at a fraction of the volume, and higher speeds
- dragonfly Delivers accurate and repeatable nanolitre to milliliter dispensing
- apricot Automated liquid handling instrumentation for convenient general use across your entire team
- Sample Preparation
-
Sample Management
- comPOUND A scalable, reliable, and secure compound management solution
- BioMicroLab Easy-to-use sample management automation instruments
- arktic Robust biospecimen storage and management down to -80°C
- lab2lab Novel sample and data transfer network system
- comPACT Reliable and efficient -20°C storage and retrieval has never been more accessible
-
Liquid Handling
-
About
- Company With a focus on liquid handling, sample preparation and sample management, our expert teams create state-of-the-art solutions that scientists and researchers can trust Culture We have one overarching mission: to work together to accelerate life science research. Through our innovative solutions and state-of-the-art tools, we believe we can make a real difference to human health Innovation From the initial prototype through to manufacturing, installation and beyond, we bring a problem-solving mindset and technical expertise to drive innovation Board Members Our Board of Directors are committed to driving the long-term success and sustainability of SPT Labtech, providing expert guidance and oversight to execute the company’s ambitious commercial strategy.
-
Positive displacement technology
.jpg) Novel positive displacement dispensing technology from disposable tips underpins our liquid handling portfolio of products
Learn more
Novel positive displacement dispensing technology from disposable tips underpins our liquid handling portfolio of products
Learn more 
-
View all
 Pneumatic tube transport
Pneumatic tube transport
.jpg) Harnessing the power of pneumatic technology, our innovative sample transport solutions deliver quickly and reliably between labs, analytical suites and even across sites hundreds of meters apart
Learn more
Harnessing the power of pneumatic technology, our innovative sample transport solutions deliver quickly and reliably between labs, analytical suites and even across sites hundreds of meters apart
Learn more 
-
Knowledge Base
- Events & Webinars Meet the SPT team at events all over the globe and virtually via our webinars Blog Our latest blog posts feature trends in research, innovative techniques and new technology Resources Our wide range of insightful resources include videos, whitepapers, eBooks, application notes and more News Latest news from SPT Labtech globally
-
23 April, 2024
 SPT Labtech supports life sciences startup growth by joining forces with BioLabs
Continue reading
SPT Labtech supports life sciences startup growth by joining forces with BioLabs
Continue reading 
-
11 April, 2024
 SPT Labtech Appoints Rob Walton as New Chief Executive Officer
Continue reading
SPT Labtech Appoints Rob Walton as New Chief Executive Officer
Continue reading 
-
27 March, 2024
 5 ways to mitigate biological sample integrity loss
Continue reading
5 ways to mitigate biological sample integrity loss
Continue reading 
10
- Careers

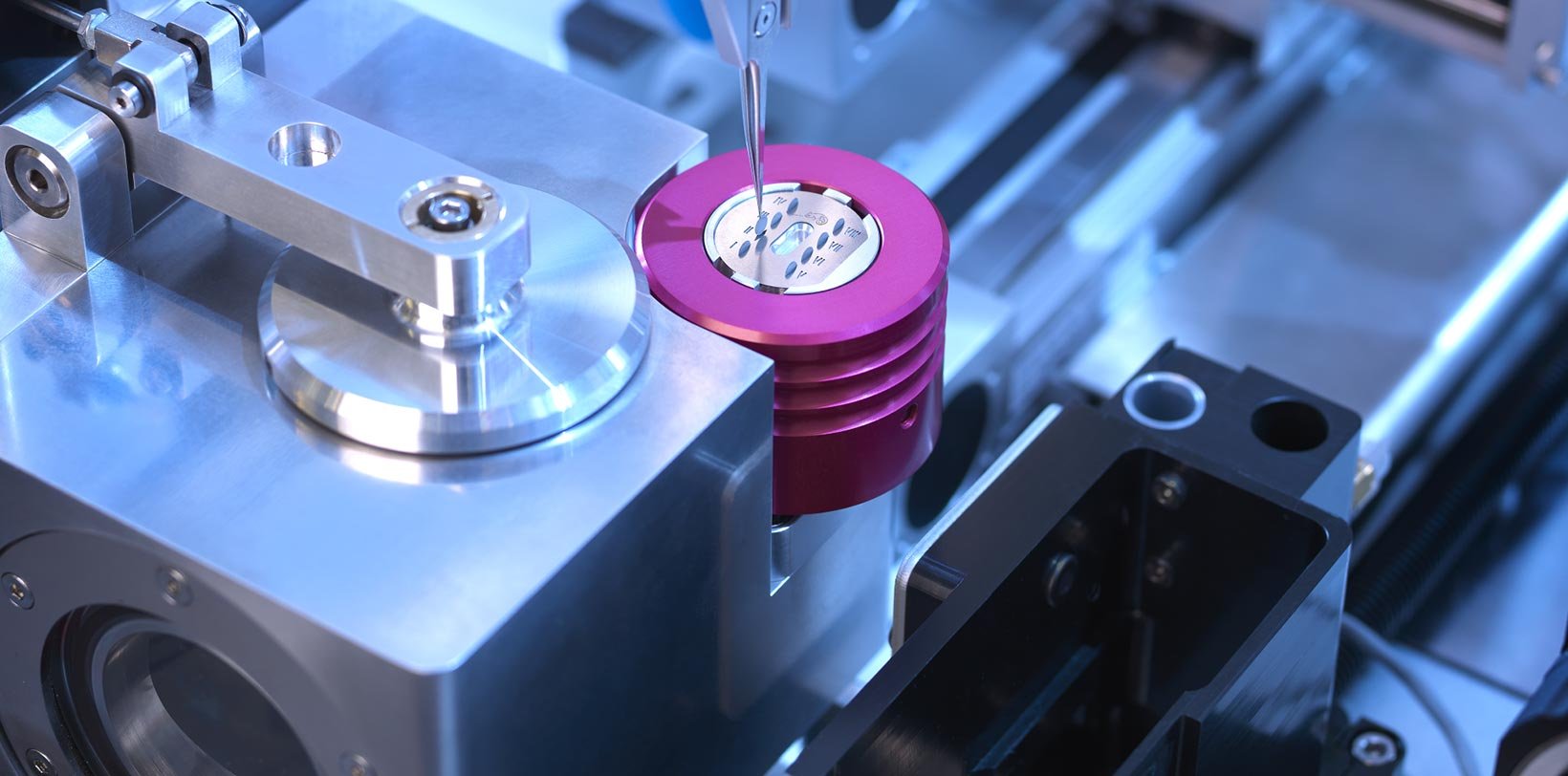
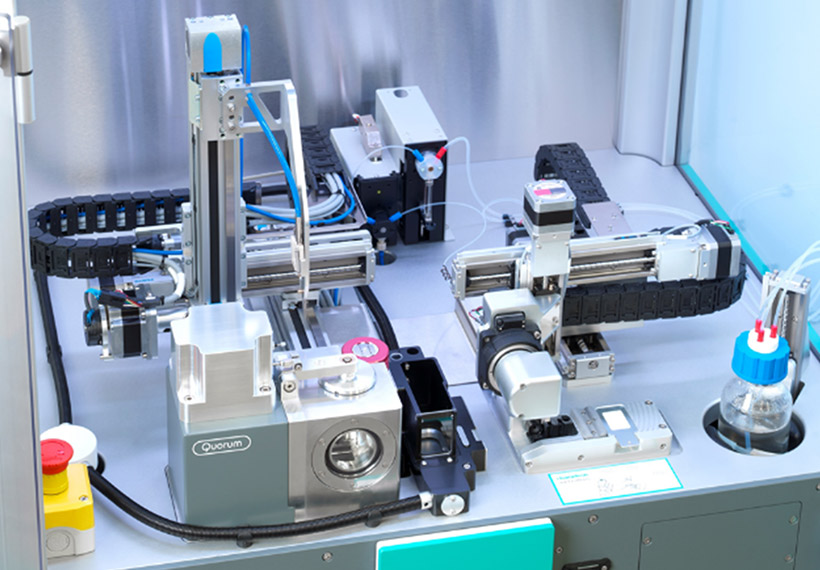
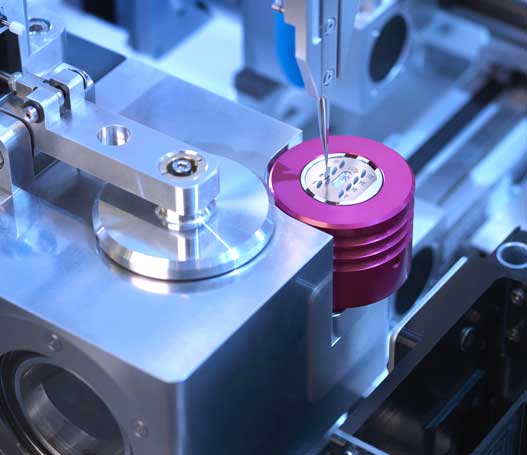
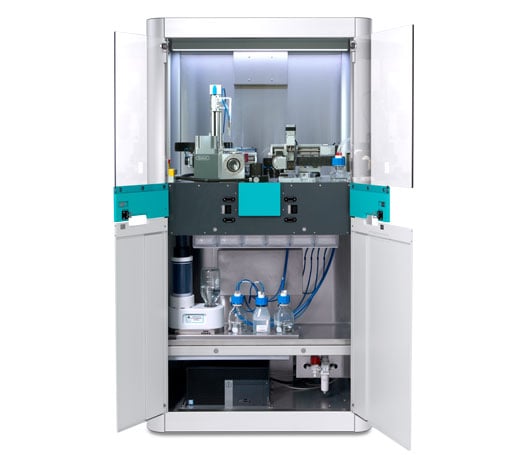
Optimize cryo-EM sample preparation workflows
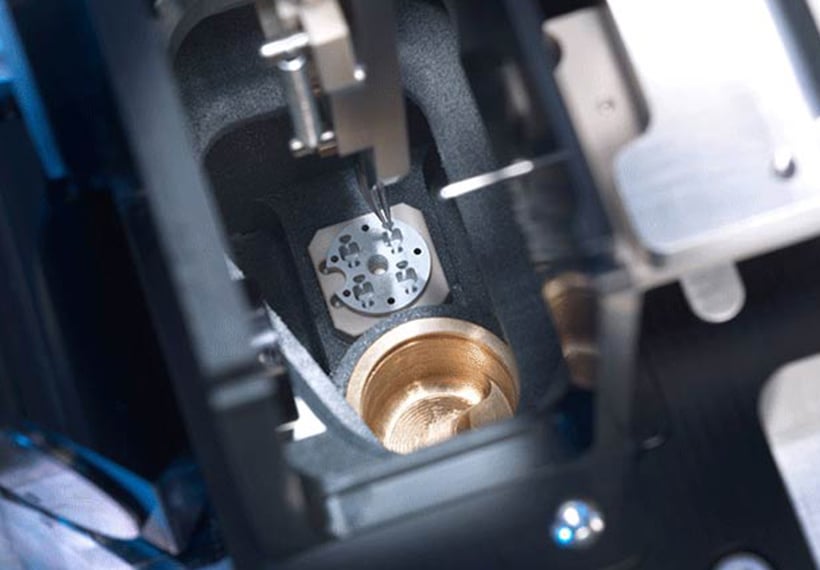
chameleon delivers optimized grid vitrification for cryo-EM by combining next-generation automation, blotless grid technology, and high speed plunging. Thinking beyond existing sample preparation workflows enables routine high resolution structural studies.
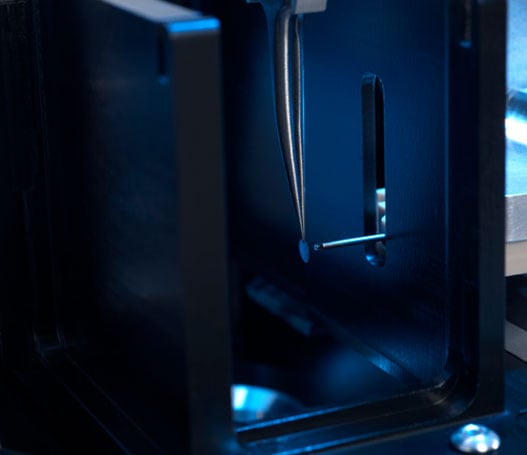
Onboard cameras provide visual images during freezing
Enabling the easier selection of only good grids based on visual images during freezing reduces the requirement for costly microscope screening and saves valuable research time.
High-speed plunging
chameleon’s high speed plunging capability reduces air-water interface protein denaturation effects and allows better handling of difficult samples to improve quality and enable better research outcomes.
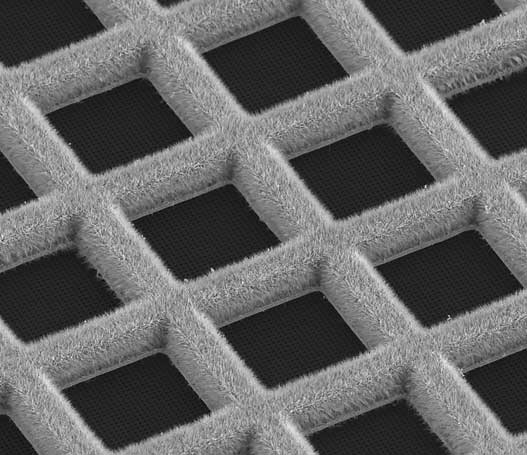
Blot-free technology
The blot-free technology using self wicking grids minimizes variability and enables researchers to gain precise sample specific control of ice thickness.
Precise automated grid handling
With almost no manual handling required, chameleon eliminates common grid damage and loss issues to increase workflow efficiency and reduce waste.
Guided workflows enable simple set up, use and cleaning
Intuitive design makes the instrument easy to use even for novice operators, minimizing training time and streamlining set up.
In-line glow discharge
Easily apply real time adjustments to perfect your sample behaviour and achieve the perfect ice thickness.
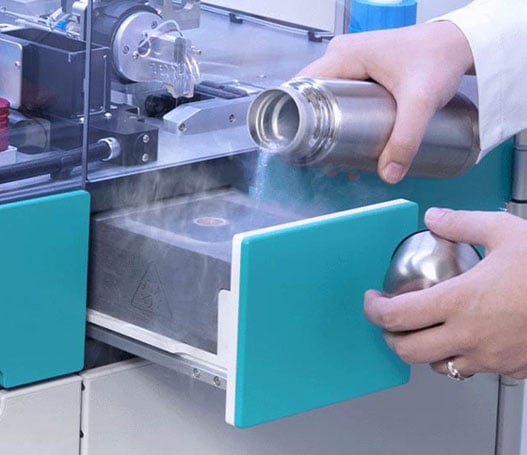
Automated cryogen level sensing and temperature control
Ensure the safe storage and control of cryogens in an automated drawer to maintain their stability and quality and assure research integrity.
Captures all relevant grid parameters and images
Improved data collection and record keeping enables future repeatability as well as data mining to inform new research avenues.





chameleon Citations
Technical Specification
Scroll to view more information
| Minimum sample volume | 5 µL | Sample block temperature control | 4°C to 37°C |
| Minimum dispense volume | 6 nL | Weight | 300kg |
| Standard dispense-to-plunge time | 101 msec | Dimensions (W x D x H) | 916 mm x 708 mm x 1687 mm |
| Fastest dispense-to-plunge time | 54 msec |

The chameleon has enabled an efficient cryoEM workflow for Arvinas by allowing us to freeze our samples promptly upon elution from the column, thus allowing rapid capture of complexes on grids ready for cryoEM data collection. By improving sample quality and reducing screening loads on microscopes, chameleon has allowed us to leverage both academic and industry partners for data collection. This allows us to focus our resources and efforts on producing high-quality protein complexes to advance the field of PROTAC® protein degradation.
Katie Digianantonio, PhD
Research Investigator









