- Applications
-
Products
-
Liquid Handling
- firefly Accelerate genomic research with innovative all-in-one, compact liquid handling
- mosquito Nanolitre liquid handling technology performs ‘traditional’ tasks at a fraction of the volume, and higher speeds
- dragonfly Delivers accurate and repeatable nanolitre to milliliter dispensing
- apricot Automated liquid handling instrumentation for convenient general use across your entire team
- Sample Preparation
-
Sample Management
- comPOUND A scalable, reliable, and secure compound management solution
- BioMicroLab Easy-to-use sample management automation instruments
- arktic Robust biospecimen storage and management down to -80°C
- lab2lab Novel sample and data transfer network system
- comPACT Reliable and efficient -20°C storage and retrieval has never been more accessible
-
Liquid Handling
-
About
- Company With a focus on liquid handling, sample preparation and sample management, our expert teams create state-of-the-art solutions that scientists and researchers can trust Culture We have one overarching mission: to work together to accelerate life science research. Through our innovative solutions and state-of-the-art tools, we believe we can make a real difference to human health Innovation From the initial prototype through to manufacturing, installation and beyond, we bring a problem-solving mindset and technical expertise to drive innovation Board Members Our Board of Directors are committed to driving the long-term success and sustainability of SPT Labtech, providing expert guidance and oversight to execute the company’s ambitious commercial strategy.
-
Positive displacement technology
.jpg) Novel positive displacement dispensing technology from disposable tips underpins our liquid handling portfolio of products
Learn more
Novel positive displacement dispensing technology from disposable tips underpins our liquid handling portfolio of products
Learn more 
-
View all
 Pneumatic tube transport
Pneumatic tube transport
.jpg) Harnessing the power of pneumatic technology, our innovative sample transport solutions deliver quickly and reliably between labs, analytical suites and even across sites hundreds of meters apart
Learn more
Harnessing the power of pneumatic technology, our innovative sample transport solutions deliver quickly and reliably between labs, analytical suites and even across sites hundreds of meters apart
Learn more 
-
Knowledge Base
- Events & Webinars Meet the SPT team at events all over the globe and virtually via our webinars Blog Our latest blog posts feature trends in research, innovative techniques and new technology Resources Our wide range of insightful resources include videos, whitepapers, eBooks, application notes and more News Latest news from SPT Labtech globally
-
23 April, 2024
 SPT Labtech supports life sciences startup growth by joining forces with BioLabs
Continue reading
SPT Labtech supports life sciences startup growth by joining forces with BioLabs
Continue reading 
-
11 April, 2024
 SPT Labtech Appoints Rob Walton as New Chief Executive Officer
Continue reading
SPT Labtech Appoints Rob Walton as New Chief Executive Officer
Continue reading 
-
27 March, 2024
 5 ways to mitigate biological sample integrity loss
Continue reading
5 ways to mitigate biological sample integrity loss
Continue reading 
10
- Careers
- Home
- Genomics
Genomics
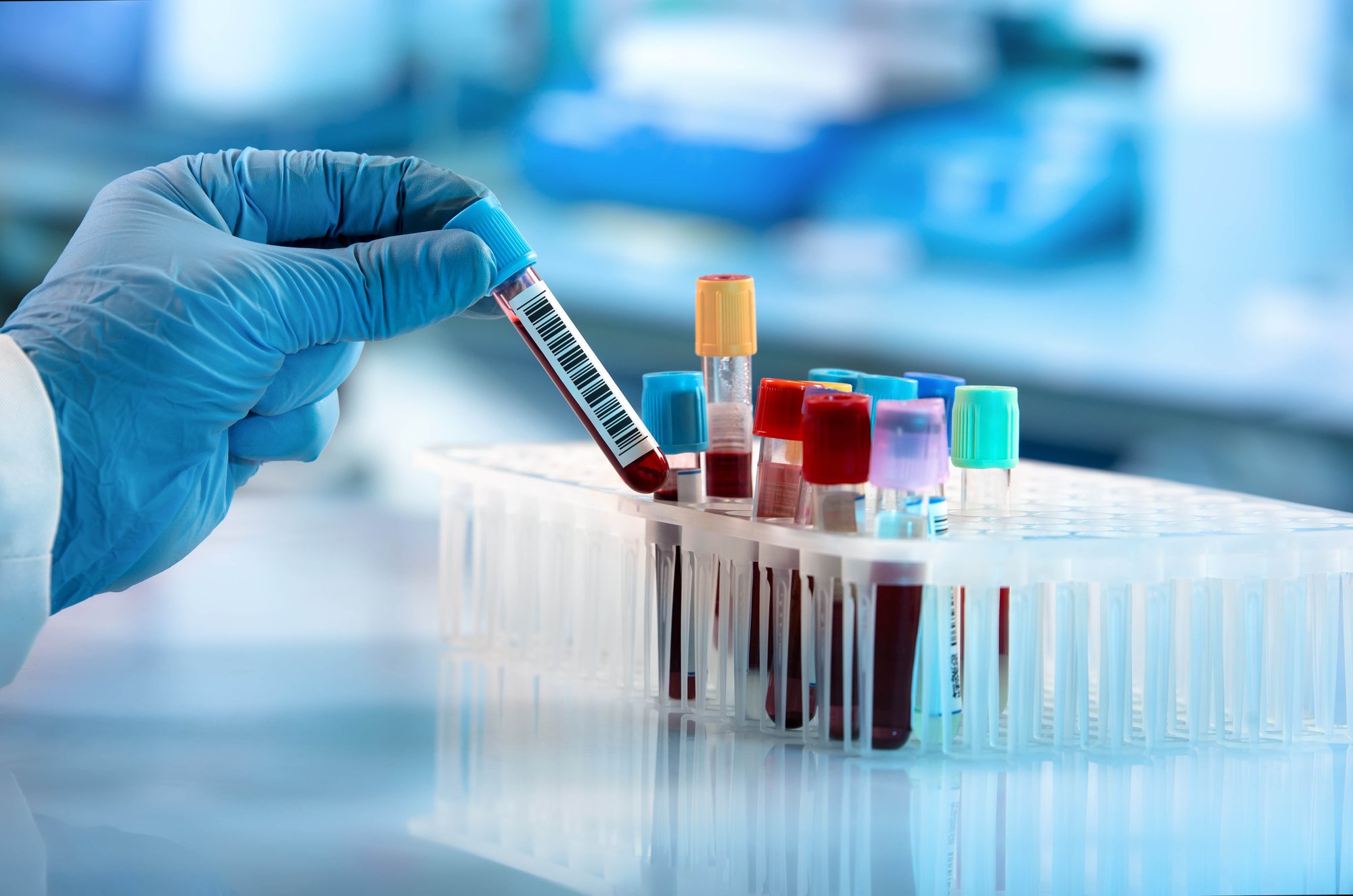
Powerful techniques from whole genome sequencing to rapidly emerging applications in areas such as spatial genomics yield rich scientific data to drive new discoveries. Laboratories are streamlining their workflows and processes to fully harness the potential of these advances.
Maximize research output in molecular biology and genomics.
An undertaking to reduce costs, or increase throughput in genomics research, has led to a demand for automation and volume miniaturization solutions to optimize sample preparation and boost research output.
Next Generation Sequencing (NGS) Library Preparation
Miniaturizing your NGS library preparation workflows dramatically reduces sample and reagent costs while enhancing the value of your data output. A wide range of kits can be miniaturized in only a few microlitres total volume has been demonstrated.
Explore NGS library preparationLaboratory Developed Tests
Laboratory Developed Tests (LDTs) are in integral tool in modern healthcare. They fulfill critical needs where existing commercial tests are not available, such as an emergency pandemic response, to further understanding into rare diseases, and to aid the development of more effective personalized cancer therapies.
Automating fundamental liquid handling tasks within LDTs boosts precision and throughput, ensuring consistently high data quality and fast turnaround times.
Find out morePCR & qPCR
Low-volume liquid handling technology enables rapid, automated setup of PCR and qPCR reactions with outstanding accuracy and precision.
Explore related productsSingle Cell Genomics
Single cell genomics provides whole genome and transcriptome sequencing from a single cell. Automation and miniaturization technologies unlock the power of single-cell analysis and sequencing making your high throughput experiments practical and cost-effective.
Explore related productsTranscriptomics
Transcriptomics uses high throughput methods to examine RNA transcripts produced by the genome. Automated nanolitre-scale liquid handling enables researchers using transcriptomics to dramatically increase throughput while balancing sensitivity and cost considerations.
Explore related productsSynthetic Biology
Synthetic biology has opened new doors for researchers now able to design large numbers of synthetic construct combinations for testing. We help you to scale up your DNA construct assembly and validation while assuring the best quality results.
Explore related productsPathogen Surveillance
Genomic-informed pathogen surveillance using high-throughput genetic sequencing is a vital tool allowing researchers to more effectively monitor infectious disease agents and positively impact public health. Pathogen surveillance researchers need robust, proven technology to deliver sample preparation at scale with miniaturization driving affordable high throughput sequencing and qPCR.
Explore related productsMicrobiome & Metagenomics
Metagenomics uncovers rich insights about microbial communities and deepens our understanding of biological systems. Metagenomics studies commonly use 16S rRNA gene sequencing, which relies on robust and precise liquid handling technology and laboratory automation solutions.
Explore related productsMeet our Field Application Scientists
Our Field Application Scientists will help you to advance your research goals throughout the life of your instrument, optimizing for the applications you need and harnessing its full potential. As specialists in a range of disciplines including genomics, the team ensures that the protocols for your applications are scientifically robust, giving you the confidence to pursue novel approaches. Working closely with customers, Field Application Scientists eliminate bottlenecks and streamline workflows to enable successful research outcomes.